Original article: Frederick Sanger, by Emanuela Pasculli
Biography
Frederick Sanger was born on 13 August 1918 in Rendcombe, England, the second of three children. The father, also named Frederick, is a doctor, a missionary, and a very religious man. The mother, Cicely, is the daughter of a rich producer of cotton. From a young age, he shows great ability in manual labor; he loves painting and woodworking. From 1932 to 1936, Sanger attends Bryanston School, where he develops a passion for biology and chemistry, and later on, he was admitted to St John’s College of Cambridge.
Despite the difficulties of his first year of university, he graduated in 1939. He decides to pursue the study of biochemistry, to which he is very passionate, enrolling in a course. In 1940 he married Joan Howe, by whom he had three children, two boys and a girl.
He is a brilliant biochemical student, so he decides to carry on with the Ph.D. achieved in 1943, with the thesis “Lysine metabolism in the animal body”. After the doctorate, he receives the ambitious award Beit Memorial Fellowship for Medical Research and from 1951 to 1983, he is a researcher at Medical Research Council.
From 1983 onwards, he withdraws from public life in a town near Cambridge and dies at 95 years old, the 19th of November 2013.
FOCUS: The invention of the Sanger sequencing method
Frederick Sanger entered Albert Chibnall’s research group, studying the amino acid structure of insulin, a hormone secreted by the pancreas, which was used to treat diabetes since 1920.
The structure of insulin and the Nobel Prize
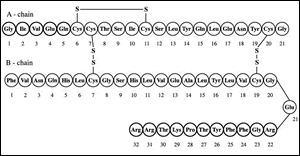
Frederick Sanger himself discovered the amino acid structure of bovine insulin in 1954 (Fig. 1). He understood that insulin consisted of two polypeptide chains held together by two disulfide bridges through chromatographic partitioning and paper chromatography. From there, he identified the amino acid of both chains: one of 21 amino acids and the other of 30. This discovery earned him the Nobel Prize in chemistry in 1958.
He kept studying the insulin structure in different mammals, comparing the results. Finally, between 1956 and 1962, he developed a procedure to determine the sequence of radioactive proteins.
The interest of nucleic acids and the second Nobel Prize
In 1958 he decided to direct his research towards molecular biology, so he became a researcher at The Medical Council Research Laboratory of Molecular Biology. He was interested in nucleic acids and, in particular, in RNA sequencing with radioactive methods. In 1967 the first RNA was sequenced, followed by a tRNA in 1969.
In 1970, the time was ripe to move to DNA sequencing. This time he developed a new method based on the acrylamide gel. This procedure, which bears his name, is used to sequence a single-stranded DNA of a virus. In 1977, Sanger and his research group sequence the phage ɸX174 (phi X174) DNA: the first genome to be sequenced (Fig. 2). Three years later, he received the second Nobel Prize. Before retiring in 1983, he sequenced the human and bovine mitochondrial genome.

The Sanger sequencing
Frederick Sanger devised a sequencing method capable of sequencing nucleic acids, from the simplest to the most complex. The method, commonly known as Sanger sequencing, is called the chain termination method.
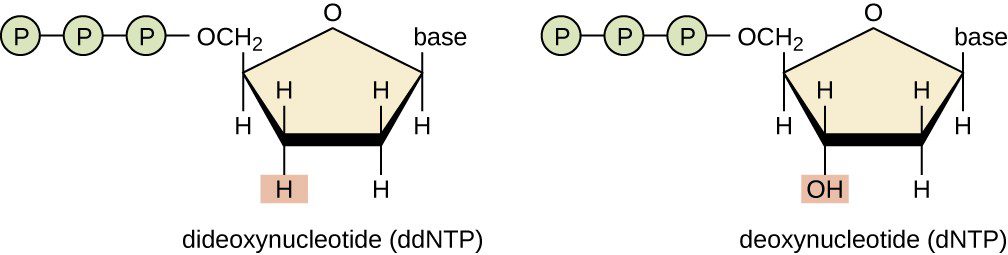
The technique consists in the employment of dideoxynucleotides (ddNTP), that compared to the deoxynucleotides (dNTP) have a modification in the sugar: they lack the hydroxyl group even at 3′ (Fig.3). This peculiar structure prevents the development of a phosphodiester bond and thus the link with another nucleotide. Therefore once they have linked a new nucleotide sequence, they block the synthesis.
The starting specimen is a denatured double-stranded DNA, so it is possible to exploit the action of DNA polymerase to synthesize the complementary strand. Four different reactions are carried out simultaneously; each has a trigger primer, a DNA polymerase, and the four dNTPs. The difference between the four reactions lies in the labeled ddNTP added – in stoichiometrically smaller amounts than the dNTP – to each reaction. In this way, the new strand, complementary to the mold, will elongate by incorporating dNTPs and will block when ddNTP is incorporated. This will create fragments of different lengths, which will be separated on a polyacrylamide gel. The nucleotidic sequence can be traced by observing the different bands formed on the gel (Fig. 4).

Scientific contribution
The Human Genome Project
The sequencing conceived by Sanger started the Genomic Era. In a few years, many goals have been reached, such as the sequencing of Haemophilus influenzae (1995) and other organisms used in research, up to the human genome sequencing (2001).
The Humane Genome Project started in 1990 as the union of different international research groups (basically from the USA, UK, Canada, and New Zealand). Finally, in 2000, there was the first draft of the genome and the complementation of the sequence after three years. This great success allows making huge steps forward in different fields, among which the biochemical one stands out. Today, in fact, since it is possible to obtain the genome of a person in a short time, it is possible to study the genetic variability, not only between individuals but also between tissues of the same person, or even between healthy cells and cancer cells.
Next Generation Sequencing (NGS)
Although the Sanger method allowed us to sequence the entire human genome, many research centers have been trying to improve this procedure. This has allowed the development of Next Generation Sequencing (NGS) methods, which groups second and third-generation methods.
The main difference between the first method is the high throughput, which allows analyzing more sequences in parallel, reducing the time of examination and costs. For the Human Genome Project, in fact, as many as 100 million dollars have been invested, compared to only 1500 dollars, required today for the sequencing of a human genome. All in a reduced time compared to the 10 years it took to obtain the first human genome.
Recognitions
Frederick Sanger is the fourth person in history to have received the Nobel Prize twice: the first time in 1958, thanks to his studies on protein structure that led him to identify the structure of bovine insulin; the second time in 1980, together with Paul Berg and Walter Gilbert, for having determined the nucleotide sequence of DNA.
Thanks to his extraordinary work in the field of genetics and biotechnology, he also received other awards:
- 1954: received the Fellowship of the Royal Society;
- 1963: became Commander of the Order of the British Empire;
- 1969: received the Royal Society’s Royal Medal;
- 1977: obtained the Copley Medal;
- 1986: became Member of the United Kingdom Order of Merit.
In 1992 the Wellcome Trust and the British Medical Research Council founded the Sanger Centre (today Sanger Institute) in his honor (Fig. 5).

Bibiography
- https://aulascienze.scuola.zanichelli.it/come-te-lo-spiego/2018/04/18/dal-metodo-sanger-a-oggi-40-anni-di-sequenziamento-del-dna/;
- Sanger, Frederick, Steven Nicklen, and Alan R. Coulson. “DNA sequencing with chain-terminating inhibitors.” Proceedings of the national academy of sciences 74.12 (1977): 5463-5467;
- https://it.wikipedia.org/;
- https://www.famousscientists.org/;
Picture bibiography
- http://www.stoccolmaaroma.it/2014/muore-frederick-sanger-due-volte-premio-nobel-per-la-chimica/
- https://www.semanticscholar.org/paper/DNA-sequencing-with-chain-terminating-inhibitors.-Sanger-Nicklen/d0bfed3a7fe8a40ddf9fba96e7b6fb1eaf5006b3
- https://www.chimicamo.org/chimica-organica/insulina/
- https://www.genomeup.com/Glossary/dideossinucleotidi/
- http://www.cusmibio.unimi.it/scaricare/Enigma_Presentation_CusMiBio.pdf
- https://www.sanger.ac.uk/about/who-we-are/sanger-institute/
Note sequencing (NGS )nel titolo le lettere sono invertite
autrice Emanuela Pasculli