Introduction
Nathan R. Smith, Francis E. Clark, and Ruth E. Gordon performed the challenge regarding the taxonomic classification of the genus Bacillus in the 1930s. Their studies allowed them to develop the definition for which the genus Bacillus is composed of “rod-shaped bacteria capable of forming endospores that are more resistant than vegetative cells to heat, desiccation, and other stressors”. A phenotypically heterogeneous group of Gram-positive organisms with an extremely broad range of nutritional requirements, growing conditions and DNA base composition characterizes the Bacillus genus. According to the definition of Ash et al.,1991, these organisms can be aerobic or facultatively anaerobic, spore-forming and rod-shaped. However, in some strains no sporulation is observed but they possess all the other properties related to sporulating strains; therefore, they have been classified as members of the genus Bacillus.
In one of the latest publications known as “Bergey’s Manual” it is possible to observe the presence of 19 different genera referable to Bacillaceae’s family such as Bacillus, Alkalibacillus, Halobacillus, Lentibacillus, Oceanobacillus, and Virgibacillus. Some species, known as part of the genus Bacillus, have been reassigned to different genera and families.
The first molecular taxonomic classification to distinguish the various domains of life is related to the use of 16s rRNA sequence. In the case of the genus Bacillus, it allowed distinguishing and precisely identify some microorganisms with sometimes overlapping phenotypic characteristics. The recent identification of species related to the Bacillus genus refers to the use of innovative methods, so algorithms, to avoid errors related to the incorrect association of species. Two algorithms are currently used: AdaptML and Ecotype Simulation, each analyze the evolutionary history of a clade to yield appropriate criteria for demarcating the significant clades corresponding to ecotypes.
These algorithms were successful for extremely closely related ecotypes that would otherwise have been considered as unrecognized organisms within a particular species. In 2018, the GTDB database listed 3407 sequences for the genus Bacillus. Considering a molecular taxonomy point of view, it is possible to obtain a more careful and accurate subdivision using the phylogenetic trees arising from the use of whole-genome sequences (Fig. 1) that allows distinguishing between closely related species rather than using 16S rRNA. This classification, based on whole-genome sequences using bioinformatic tools, provide a step-change in reliability, as evidenced by significantly high average bootstrap in phylogenomic trees.
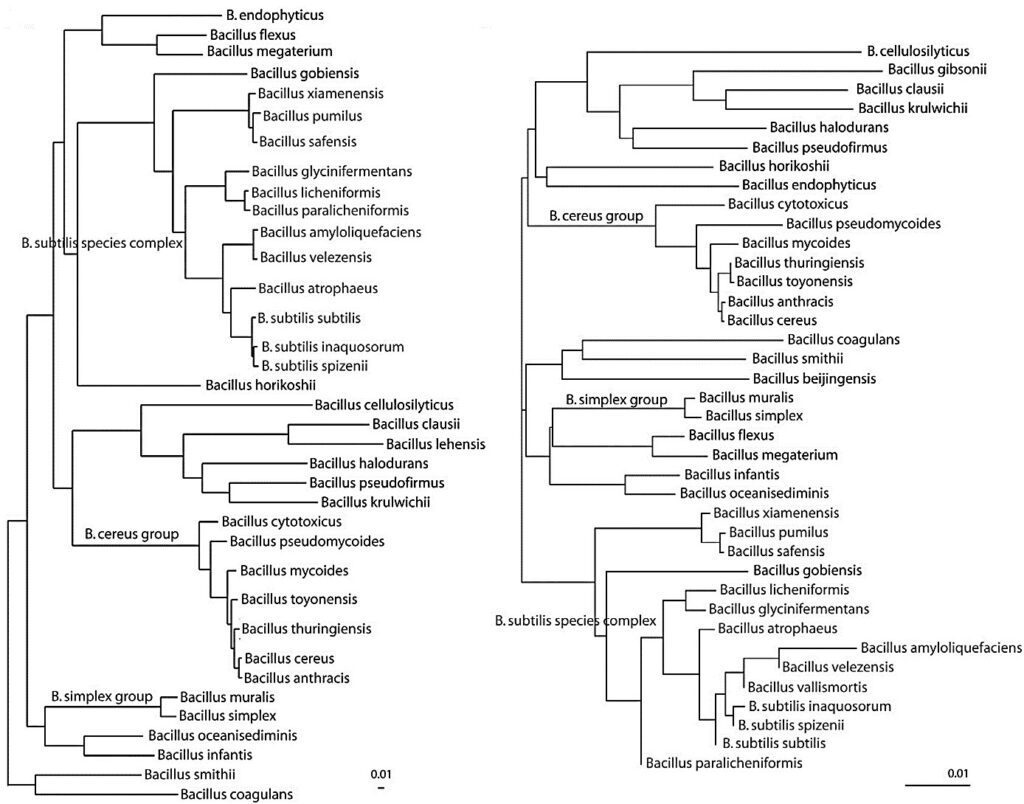
Simultaneously, DNA analysis also have pushed estimates towards millions of existing species. A corresponding method is Average Nucleotide Identity (ANI). The method is based on the pairwise calculation of nucleotide identity between core genomes.
According to the initial classification realized in the 1930s, the use of 16s rRNA sequencing and the latest classifications, it is possible to distinguish separation of the genus Bacillus into two groups: “Bacillus sensu lato”, consisting of bacteria capable of forming endospores, and “Bacillus sensu sticto” which represents the group consisting of the original species of bacillus. Lastly, among the original species of the genus Bacillus we recognize Bacillus subtilis which represents the species with the oldest nomenclature.
Identification methods of Bacillus
Identification studies based on 16s rRNA sequencing are often used to include or exclude an organism concerning a particular genus but is not possible to realize an accurate identification between species that are closely related. For this reason, it is not possible to consider this approach as a useful tool for the identification of new species but only for a basic phylogenetic analysis. One of the techniques useful for precise identification is DNA-DNA cross-hybridization that is often associated with the comparison of G-C % content within the genome. However, there are a various number of other useful methods as shown in the following table reported in one of the recent papers written by Goldman et al., 2009.
Table 1. Common methods used for the identification of the genus Bacillus.
| Method | Useful range |
| Physiological and morphological tests | |
| Microscopy and 26-test battery, Microscopy and 20-test battery, Carbohydrate utilization | Species |
| DNA sequencing | |
| Single locus sequence analysis: 16S rRNA and 23S rRNA – groEL, gyrB, recN, rpoB, spoIIA | Domain-genus/Species-subspecies |
| Multilocus Sequence Typing (MLST): adk, ccpA, ftsA, glpT, pyrE, recF, sucC, glpF, gmk, ilvD, pta, pur, pycA, tpi, rpoB, gyrB, pycA, mdh, mbl, mutS, plcR | Genus-subspecies |
| DNA fingerprinting | |
| Restriction Fragment Length Polymorphism (RFLP): rRNA operons, amplified secY and gyrB genes | Subspecies-strain/Genus-species |
| Pulsed Field Gel Electrophoresis (PFGE), Repetitive DNA PCR fingerprinting, Random Amplified Polymorphic DNA (RAPD), Oligonucleotide microarray fingerprinting | Subspecies-strain |
| Other molecular methods | |
| Fatty acid profiling, MALDI-TOF mass spectroscopy of spore proteins | Species-subspecies |
| Multilocus Enzyme Electrophoresis (MLEE) | Subspecies-strain |
The selection of the useful method for the identification is closely related to the preliminary information possessed and to the final objective to be achieved as the inclusion within a pre-existing group or the distinction of microorganism as a new species not linked to any group. Therefore, the work of taxonomic identification and organization represents an ever-open challenge considering the number of species that are continually discovered. Besides, systematic studies are needed to give a greater value to this genre and highlight new utilities.
Industrial and medical applications of Bacillus
Some products, based on Bacillus spores, are suitable for industrial products because can be alive for nearly unlimited time and can be stored like chemical products. Bacillus spp. are highly adapted to abiotic and biotic stress. Considering this aspect, Bacillus species play a key role within the modern industry. The use of metabolic products belonging to these species makes it possible to reduce costs and replace old chemical processing processes with much more environmentally friendly ones.
There are various types of commercial products that can be obtained such as antibiotics, vitamins, and food enzymes. Moreover, due to their plant growth-promoting activity, many species of Bacillus are commercially used as phytostimulators and biofertilizers. Biocontrol products based on Bacillus spp. are broadly applied in agriculture as valid substitutes of chemical pesticides: some strains of Brevibacillus have a well-known larvicidal activity, can be used as biological control of plant pathogens and can improve the shelf-life of fruits.
In the environmental context, in addition, it is possible to identify some species of Bacillus able to act as bioremediators in fact, Lysinibacillus species show the ability to degrade harmful insecticides, selectively desulfurize dibenzothiophene and create different types of larvicidal toxins. The general products obtained by the action of Bacillus microorganisms are often considered as safe according to their natural origin, in fact, in the United States these products are recognized as GRAS (Generally Regarded As Safe) while in the European Union the acronym QPS (Qualified Presumption of Safety) has been proposed. The wide variety of uses is related both to the excellent natural competence that these organisms can develop and to their ability to activate DNA recombination mechanisms.
New tools have been developed and tested primarily on B. subtilis, and these include so-called plasmid cloning vectors, transposon mutagenesis systems, antibiotic resistance markers and single- and multiple- copy gene expression cassettes associated with the production of high-value proteins and enzymes. Enzymes that are produced, used, or marketed can be detergents, starch hydrolysis, textiles, baking and beverages. These products have special features, in fact, can resist high temperatures, alkaline conditions or wide pH ranges.
In the context of vitamins production, instead, is possible to find the presence of riboflavin, poly-γ-glutamic acid, D-ribose and polyhydroxy butyrate (PHB). Within modern medicine, there is various evidence that shows how microorganisms can affect the health status of individuals. Some species of Bacillus, such as B. anthracis, are particularly known for their ability to create health problems and sometimes can even be used for warfare purposes considering the hazardous topic of bioterrorism. However, many species have a largely positive impact on the health of individuals thanks to the natural production of antibiotic substances that can be used during hospital treatment.
In addition to antibiotic substances, some Bacillus species are helpful to produce human proteins and vaccines. These processes have been made using B. subtilis, B. megaterium, and B. brevis. Other applications include prophylactic use and as health food supplements. In this last case, the species found in the products are B. subtilis, B. licheniformis, B. coagulans, B. pumilus, and B. clausii. Also, these same microorganisms are considered very powerful animal probiotics, especially in the pig/poultry sectors and in aquaculture.
Based on the various functionalities that this group of microorganisms possesses, the high potential for biotechnological application and the ability to withstand adverse conditions, it is possible to consider Bacillus as the best organism to be used in industrial production and quality control processes. Considering B. subtilis, the particular survival and resistance characteristics of its spores, is possible to consider an innovative use of pre‐calibrated biofilms with spores as biological UV‐dosimeters in space during several space missions.
Another application in which the use of microbes can be useful is the process called microbial enhanced oil recovery (MEOR), which is used to recover crude oil from “depleted” reservoirs. The Bacillus species are used in these processes for their ability to produce surfactin and exopolysaccharide. Concerning the extraordinary applications proposed, it is important to emphasize that the data collection about its genetics, spore resistance/germination and metabolites produced is a powerful tool to obtain an increasingly accurate assessment of the positive impact that some microorganisms can have on the scientific development, economy and world health.
References
- Ash, C., J.A.E. Farrow, S. Wallbanks, and M.D. Collins. 1991. Phylogenetic heterogeneity of the genus Bacillus revealed by comparative analysis of small-subunit-ribosomal RNA sequences. Lett. Appl. Microbiol.,13:202–206
- Ashiuchi, M., K. Shimanouchi, T. Horiuchi, T. Kamei, and H. Misono. 2006. Genetically engineered poly-gamma-glutamate producer from Bacillus subtilis ISW1214. Biosci. Biotechnol. Biochem., 70:1794–1797
- Bahuguna, A., M. K. Lily, A. Munjal, R. N. Singh, and K. Dangwal, 2011. Desulfurization of dibenzothiophene (dbt) by a novel strain lysinibacillus sphaericus dmt-7 isolated from diesel contaminated soil. Journal of Environmental Sciences 23: 975–982
- Banat, I. M., A. Franzetti, I. Gandolfi, et al. 2010. Microbial biosurfactants production, applications and future potential. Appl. Microbiol. Biotechnol. 87:427‐444
- Borriss, R. 2020. Bacillus. Beneficial Microbes in Agro-Ecology, 107–132. DOI: 10.1016/b978-0-12-823414-3.00007-1
- Chandel, S., E. J. Allan, and S.Woodward, 2010. Biological control of fusarium oxysporum f. sp. lycopersici on tomato by brevibacillus brevis. Journal of Phytopathology 158: 470–478
- Che, J.-m., X.-f. Zheng, B. Liu, M.-x. Su, and Y.-j. Zhu, 2011. Preparation of brevibacillus brevis fjat-0809-glx angent and study on its effect on loquats (eriobotrya japonica). Storage & Process 5
- Choi, Y.-J., T.-T. Wang, and B.H. Lee. 2002. Positive selection vectors. Crit. Rev. Biotechnol., 22:225–244
- Conn, H. J. 1930. The identity of Bacillus subtilis. The Journal of Infectious Diseases, 341-350
- Connor, N., Sikorski, J., Rooney, A. P., Kopac, S., Koeppel, A. F., Burger, A., Cohan, F. M. 2010. Ecology of Speciation in the Genus Bacillus. Applied and Environmental Microbiology, 76(5), 1349–1358. DOI: 10.1128/aem.01988-09
- Dale, G.E., H. Langen, M.G. Page, R.L. Then, and D. Stuber. 1995. Cloning and characterization of a novel, plasmid-encoded trimethoprim-resistant dihydrofolate reductase from Staphylococcus haemolyticus MUR313. Antimicrob. Agents Chemother., 39:1920–1924
- Goldman E., H Green L., 2009. Practical handbook of microbiology. Second edition. CRC Press. Taylor & Francis Group. ISBN 978-0-8493-9365-5
- Gordon, R.E., W.C. Haynes, and C.H.-N. Pang. 1973. The genus Bacillus. Washington, DC: U.S. Government Printing Office
- Gordon, Benjamin & Duellman, Paul & Salvucci, Anthony & Lorme, Marthah. 2019. Detecting dipicolinic acid production and biosynthesis pathways in Bacilli and Clostridia. DOI: 10.1101/803486
- Gudina, E. J., J. F. Pereira, R. Costa, J. A. Coutinho, J. A. Teixeira, and L. R. Rodrigues. 2013. Biosurfactantproducing and oil‐degrading Bacillus subtilis strains enhance oil recovery in laboratory sand‐pack columns. J. Hazard. Mater. 261:106‐113
- Hamoen, L.W., G. Venema, and O.P. Kuipers. 2003. Controlling competence in Bacillus subtilis: shared use of regulators. Microbiology, 149:9–17
- Horneck, G., D. M. Klaus, and R. L. Mancinelli. 2010. Space microbiology. Microbiol. Mol. Biol. Rev. 74:121‐156
- Le Breton, Y., N.P. Mohapatra, and W.G. Haldenwang. 2006. In vivo random mutagenesis of Bacillus subtilis by use of TnYLB-1, a mariner-based transposon. Appl. Envir. Microbiol., 72:327–333
- Lozano, L. C., J. A. Ayala, and J. Dussán. 2011. Lysinibacillus sphaericus s-layer protein toxicity against culex quinquefasciatus. Biotechnology Letters 33: 2037–2041
- Meima, R.B., J.M. van Dijl, S. Holsappel, and S. Bron. 2004. Expression systems in Bacillus. In Protein Expression Technologies: Current Status and Future Trend
- Rettberg, P., and G. Horneck. 2000. Biologically weighted measurement of UV radiation in space and on Earth with the biofilm technique. Adv. Space Res. 26:2005‐2014
- Richter, M., Rosselló-Móra, R., Glöckner, F.O., Peplies, J., 2016. JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32, 929e931. DOI: https://doi.org/10.1093/bioinformatics/btv681
- Schneider, K.R., M.E. Parish, R.M. Goodrich, and T. Cookingham. 2004. Preventing foodborne illness: Bacillus cereus and Bacillus anthracis, University of Florida (FSHN04-05 http://edis.ifas.ufl.edu/FS103)
- Sen, R. 2008. Biotechnology in petroleum recovery: The microbial EOR. Progress in Energy and Combustion Science, 34(6), 714–724. DOI: 10.1016/j.pecs.2008.05.001
- Singh, B., J. Kaur, and K. Singh, 2012. Transformation of malathion by lysinibacillus sp. isolated from soil. Biotechnology Letters 34:22 863–867
- Stackebrandt, E., W. Frederiksen, G.M. Garrity, et al. 2002. Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology. Int. J. Syst. Evol. Microbiol., 52:1043–1047
- Stein, T. 2005. Bacillus subtilis antibiotics: structures, syntheses and specific functions. Mol. Microbiol., 56:845–857
- Tam, N.K.M., N. Q. Uyen, H.A. Hong, et al. 2006. The intestinal life cycle of Bacillus subtilis and close relatives. J. Bacteriol., 188:2692–2700
- Terpe, K. 2006. Overview of bacterial expression systems for heterologous protein production: from molecular and biochemical fundamentals to commercial systems. Appl. Microbiol. Biotechnol., 72:211–222
- Zeigler, D.R. 2005. Application of a recN sequence similarity analysis to the identification of species within the bacterial genus Geobacillus. Int. J. Syst. Evol. Microbiol., 55:1171–1179
- Zubasheva, M., L. Ganushkina, T. Smirnova, and R. Azizbekyan, 2010. Larvicidal activity of crystal-forming strains of brevibacillus laterosporus. Applied Biochemistry and Microbiology 46:755–762
- Donna vettore creata da macrovector – it.freepik.com